mlens
ML-Ensemble combines a Scikit-learn high-level API with a low-level computational graph framework to build memory efficient, maximally parallelized ensemble networks in as few lines of codes as possible.
ML-Ensemble is thread safe as long as base learners are and can fall back on memory mapped multiprocessing for memory-neutral process-based concurrency. For tutorials and full documentation, visit the project website.
Ensembles as computational graphs
An ensemble is built on top of a computational graph,
allowing users great design freedom. Ensembles can be built with recursion,
dynamic evaluation (e.g. if-else) and much more. A high-level API wraps common
ensemble architectures into Scikit-learn estimators.
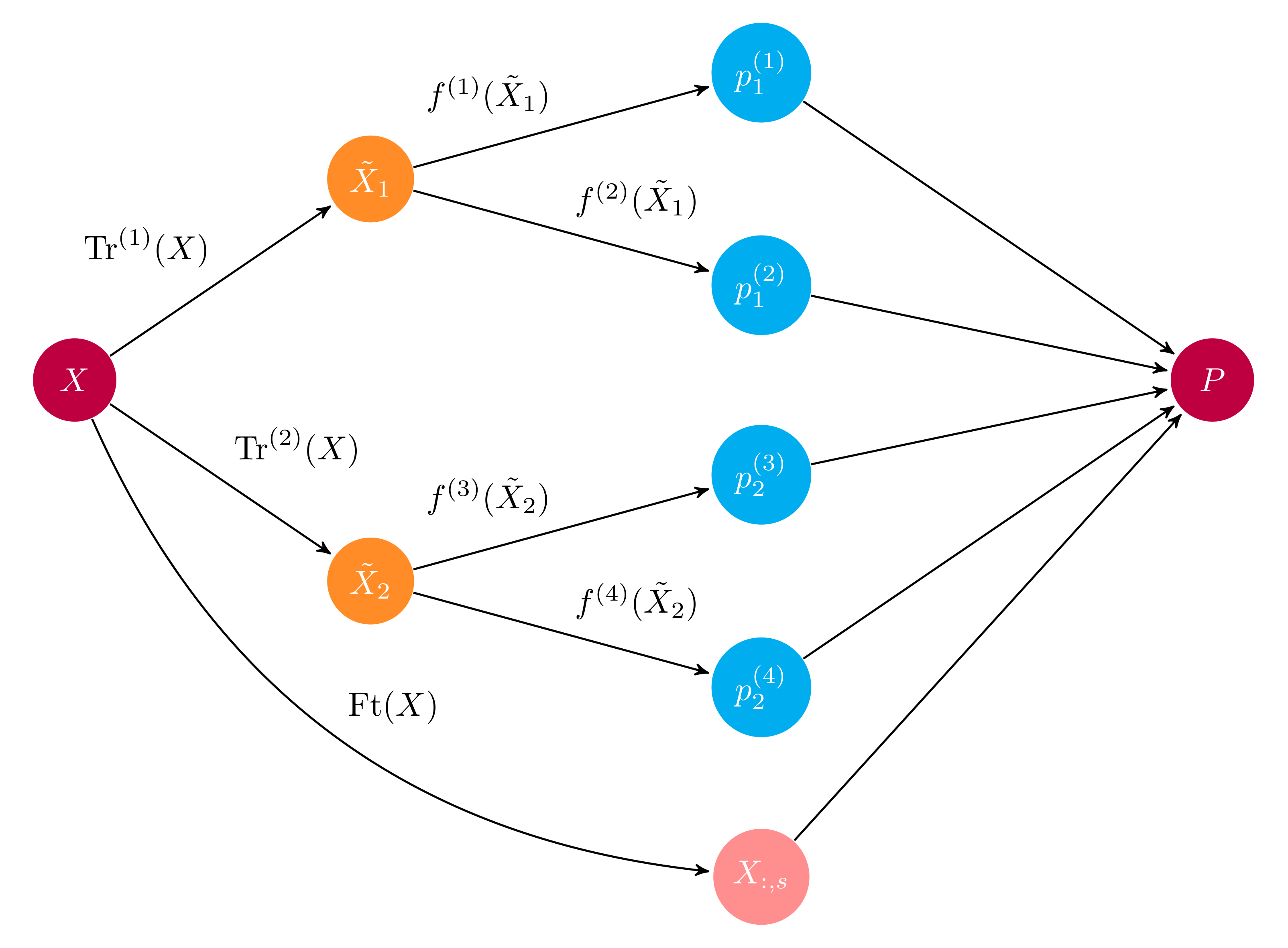
Example computational graph of a layer in an ensemble
Memory efficient parallelized learning
ML-Ensemble is optimized for speed and minimal memory consumption. No
serialization of data takes place, regardless of whether multithreading or
multiprocessing is used. Additionally, multithreading is pickle-free.
Easy of use
Ready-made ensembles are built by adding layers to an instance. No matter how
complex the ensemble, to train it call the fit method:
ensemble = Subsemble()
# First layer
ensemble.add(list_of_estimators)
# Second layer
ensemble.add(list_of_estimators)
# Final meta estimator
ensemble.add_meta(estimator)
# Train ensemble
ensemble.fit(X, y)
Similarly, it's straightforward to modify an existing ensemble:
# Remove layer
ensemble.remove(2)
# Change layer parameters
ensemble.replace(0, new_list_of_estimators)
And to create differentiated preprocessing pipelines for different subsets
of estimators within a given layer, simple pass a mapping to the add
method:
preprocessing = {'pipeline-1': list_of_transformers_1,
'pipeline-2': list_of_transformers_2}
estimators = {'pipeline-1': list_of_estimators_1,
'pipeline-2': list_of_estimators_2}
ensemble.add(estimators, preprocessing)
Dedicated diagnostics
ML Ensemble implements a dedicated diagnostics and model selection suite
for intuitive and speedy ensemble evaluation. The Evaluator
allows you to evaluate several preprocessing pipelines and several
estimators in one go, giving you birds-eye view of how different candidates
for the ensemble perform.
Moreover, entire ensembles can be used as a preprocessing pipeline, to
leverage model selection for higher-level layers. Simply set model_selection
to True in the ensemble (don't forget to turn it off when done).
preprocessing_dict = {'pipeline-1': list_of_transformers_1,
'pipeline-2': list_of_transformers_2}
evaluator = Evaluator(scorer=score_func)
evaluator.fit(X, y, list_of_estimators, param_dists_dict, preprocessing_dict)
All ensembles and model selection instances provides summary statistics in
tabular format. You can find fit and prediction times in any ensemble by calling
data, and cv-scores if you passed a scorer. For the model selection suite,
the results attribute gives you the outcome of an evaluation::
fit_time_mean fit_time_std train_score_mean train_score_std test_score_mean test_score_std params
prep-1 est-1 0.001353 0.001316 0.957037 0.005543 0.960000 0.032660 {}
est-2 0.000447 0.000012 0.980000 0.004743 0.966667 0.033333 {'n_neighbors': 15}
prep-2 est-1 0.001000 0.000603 0.957037 0.005543 0.960000 0.032660 {}
est-2 0.000448 0.000036 0.965185 0.003395 0.960000 0.044222 {'n_neighbors': 8}
prep-3 est-1 0.000735 0.000248 0.791111 0.019821 0.780000 0.133500 {}
est-2 0.000462 0.000143 0.837037 0.014815 0.800000 0.126491 {'n_neighbors': 9}
Install
PyPI
ML-Ensemble is available on PyPI. Install with
pip install mlens
Bleeding edge
Fork the GitHub repository:
git clone https://github.com/flennerhag/mlens.git; cd mlens;
python install setup.py
Citation
For scientific publication, ML-Ensemble can be cited as
@misc{flennerhag:2017mlens,
author = {Flennerhag, Sebastian},
title = {ML-Ensemble},
month = nov,
year = 2017,
doi = {10.5281/zenodo.1042144},
url = {https://dx.doi.org/10.5281/zenodo.1042144}
}



