pae_to_domains
Graph-based community clustering approach to extract protein domains from a predicted aligned error matrix
Overview
Using a predicted aligned error matrix corresponding to an AlphaFold2 model (e.g. as downloaded from https://alphafold.ebi.ac.uk/), returns a series of lists of residue indices, where each list corresponds to a set of residues clustering together into a pseudo-rigid domain.
Requirements
- Python >=3.7
- NetworkX >= 2.6.2
Known Issues
Due to an internal implementation issue in NetworkX (Issue #4992) some combinations of PAE matrix and resolution can lead to a KeyError. Solutions to this are being explored, and it will hopefully be fixed in the next NetworkX release.
Usage
While primarily intended as a code snippet to be incorporated into larger projects, this can also be called from the command line. At its simplest:
python pae_to_domains.py pae_file.json
... will yield a .csv file with each line providing the indices for one residue cluster. Full help for the command-line version:
positional arguments:
pae_file Name of the PAE JSON file.
optional arguments:
-h, --help show this help message and exit
--output_file OUTPUT_FILE
Name of output file (comma-delimited text format.
Default: clusters.csv
--pae_power PAE_POWER
Graph edges will be weighted as 1/pae**pae_power.
Default: 1.0
--pae_cutoff PAE_CUTOFF
Graph edges will only be created for residue pairs
with pae<pae_cutoff. Default: 5.0
--resolution RESOLUTION
Higher values lead to stricter (i.e. smaller)
clusters. Default: 1.0
Example
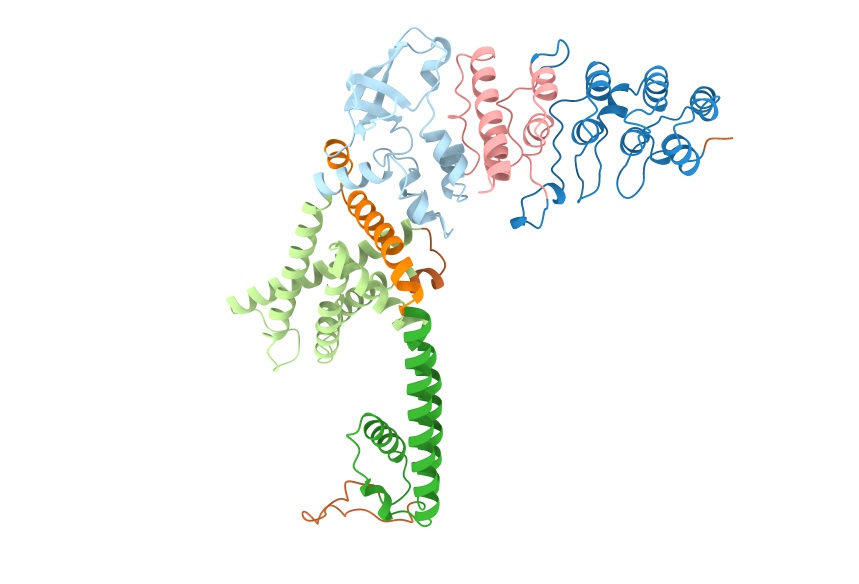
Using https://alphafold.ebi.ac.uk/entry/Q9HBA0 as an example case...
resolution=0.5:
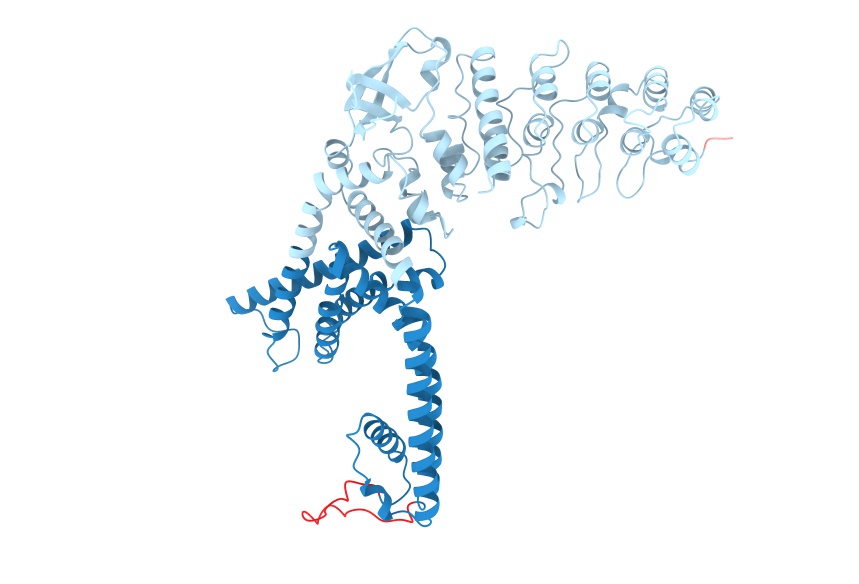
resolution=1.0:
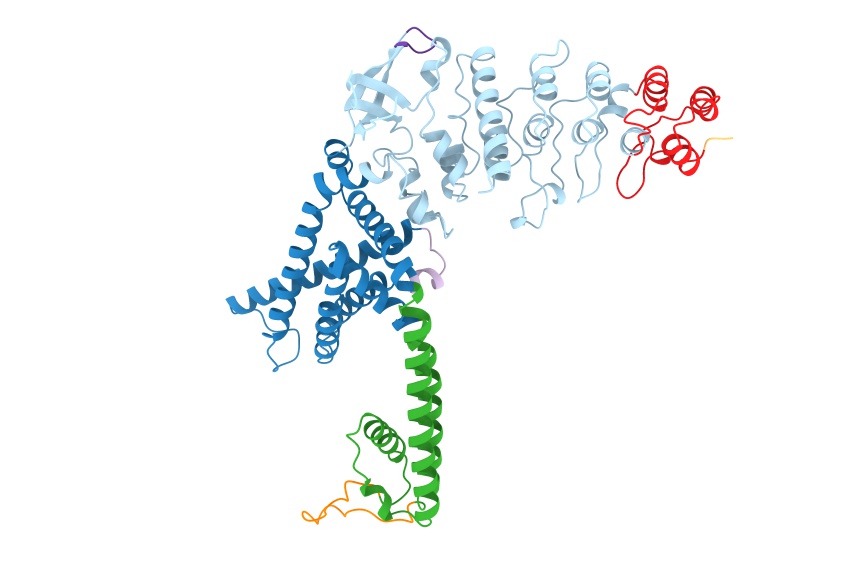
resolution=2.0: