Splitter
A PyTorch implementation of Splitter: Learning Node Representations that Capture Multiple Social Contexts (WWW 2019).
Abstract
Recent interest in graph embedding methods has focused on learning a single representation for each node in the graph. But can nodes really be best described by a single vector representation? In this work, we propose a method for learning multiple representations of the nodes in a graph (e.g., the users of a social network). Based on a principled decomposition of the ego-network, each representation encodes the role of the node in a different local community in which the nodes participate. These representations allow for improved reconstruction of the nuanced relationships that occur in the graph a phenomenon that we illustrate through state-of-the-art results on link prediction tasks on a variety of graphs, reducing the error by up to 90%. In addition, we show that these embeddings allow for effective visual analysis of the learned community structure.
This repository provides a PyTorch implementation of Splitter as described in the paper:
Splitter: Learning Node Representations that Capture Multiple Social Contexts.
Alessandro Epasto and Bryan Perozzi.
WWW, 2019.
[Paper]
The original Tensorflow implementation is available [here].
Requirements
The codebase is implemented in Python 3.5.2. package versions used for development are just below.
networkx 1.11
tqdm 4.28.1
numpy 1.15.4
pandas 0.23.4
texttable 1.5.0
scipy 1.1.0
argparse 1.1.0
torch 1.1.0
gensim 3.6.0
Datasets
The code takes the **edge list** of the graph in a csv file. Every row indicates an edge between two nodes separated by a comma. The first row is a header. Nodes should be indexed starting with 0. A sample graph for `Cora` is included in the `input/` directory.
Outputs
The embeddings are saved in the `input/` directory. Each embedding has a header and a column with the node IDs. Finally, the node embedding is sorted by the node ID column.
Options
The training of a Splitter embedding is handled by the `src/main.py` script which provides the following command line arguments.
Input and output options
--edge-path STR Edge list csv. Default is `input/chameleon_edges.csv`.
--embedding-output-path STR Embedding output csv. Default is `output/chameleon_embedding.csv`.
--persona-output-path STR Persona mapping JSON. Default is `output/chameleon_personas.json`.
Model options
--seed INT Random seed. Default is 42.
--number of walks INT Number of random walks per node. Default is 10.
--window-size INT Skip-gram window size. Default is 5.
--negative-samples INT Number of negative samples. Default is 5.
--walk-length INT Random walk length. Default is 40.
--lambd FLOAT Regularization parameter. Default is 0.1
--dimensions INT Number of embedding dimensions. Default is 128.
--workers INT Number of cores for pre-training. Default is 4.
--learning-rate FLOAT SGD learning rate. Default is 0.025
Examples
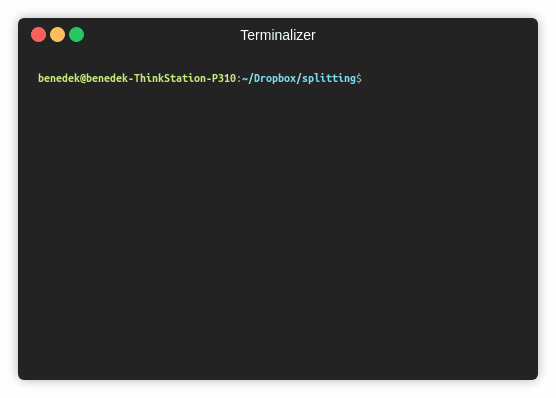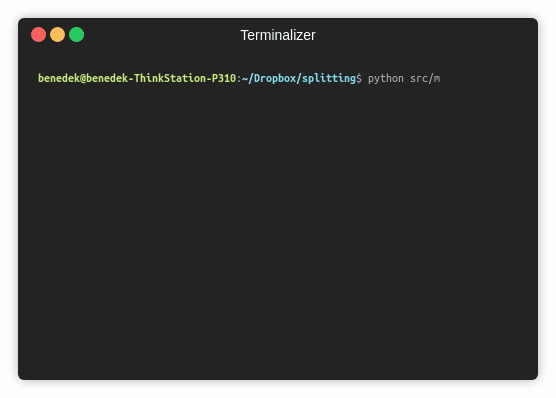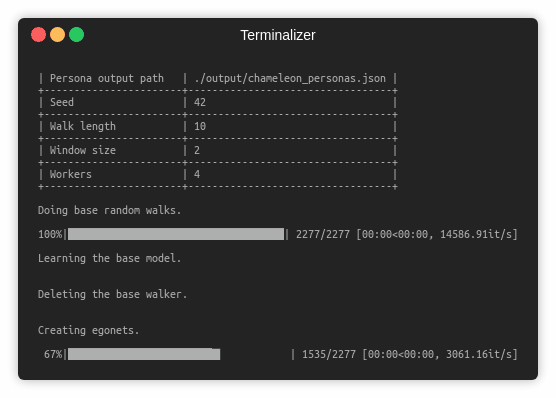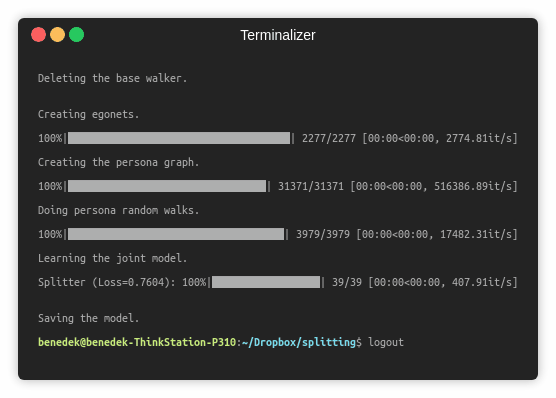
The following commands learn an embedding and save it with the persona map. Training a model on the default dataset.
python src/main.py
Training a Splitter model with 32 dimensions.
python src/main.py --dimensions 32
Increasing the number of walks and the walk length.
python src/main.py --number-of-walks 20 --walk-length 80







