TorchIO
including intensity and spatial transforms for data augmentation and preprocessing. Transforms include typical computer vision operations such as random affine transformations and also domain specific ones such as simulation of intensity artifacts due to MRI magnetic field inhomogeneity or k-space motion artifacts.
This package has been greatly inspired by NiftyNet.
Installation
$ pip install torchio
Features
Data handling
ImagesDataset
ImagesDataset is a reader of medical images that directly inherits from
torch.utils.Dataset.
It can be used with a
torch.utils.DataLoader
for efficient reading and data augmentation.
The paths suffix must be .nii, .nii.gz or .nrrd.
import torchio
subject_a = {
't1': dict(path='~/Dropbox/MRI/t1.nii.gz', type=torchio.INTENSITY),
'label': dict(path='~/Dropbox/MRI/t1_seg.nii.gz', type=torchio.LABEL),
}
subject_b = {
't1': dict(path='/tmp/colin27_t1_tal_lin.nii.gz', type=torchio.INTENSITY),
'label': dict(path='/tmp/colin27_seg1.nii.gz', type=torchio.LABEL),
}
paths_list = [subject_a, subject_b]
subjects_dataset = torchio.ImagesDataset(paths_list)
subject_sample = subjects_dataset[0]
Samplers
torchio includes grid, uniform and label patch samplers. There is also an
aggregator used for dense predictions. The code for these is almost
copy-pasted from NiftyNet.
For more information about patch-based training, see
NiftyNet docs.
import torch
import torchio
CHANNELS_DIMENSION = 1
patch_overlap = 4
grid_sampler = torchio.inference.GridSampler(
input_array, # some NumPy array
patch_size=128,
patch_overlap=patch_overlap,
)
patch_loader = torch.utils.data.DataLoader(grid_sampler, batch_size=4)
aggregator = torchio.inference.GridAggregator(
input_array,
patch_overlap=patch_overlap,
)
with torch.no_grad():
for patches_batch in patch_loader:
input_tensor = patches_batch['image']
locations = patches_batch['location']
logits = model(input_tensor) # some torch.nn.Module
labels = logits.argmax(dim=CHANNELS_DIMENSION, keepdim=True)
outputs = labels
aggregator.add_batch(outputs, locations)
output_array = aggregator.output_array
Queue
A patches Queue (or buffer) can be used for randomized patch-based sampling
during training.
This interactive animation
can be used to understand how the queue works.
import torch
import torchio
patches_queue = torchio.Queue(
subjects_dataset=subjects_dataset,
queue_length=300,
samples_per_volume=10,
patch_size=96,
sampler_class=torchio.sampler.ImageSampler,
num_workers=4,
shuffle_subjects=True,
shuffle_patches=True,
)
patches_loader = DataLoader(patches_queue, batch_size=4)
num_epochs = 20
for epoch_index in range(num_epochs):
for patches_batch in patches_loader:
logits = model(patches_batch) # model is some torch.nn.Module
Transforms
The transforms package should remind users of
torchvision.transforms.
They take as input the samples generated by an
ImagesDataset.
Intensity
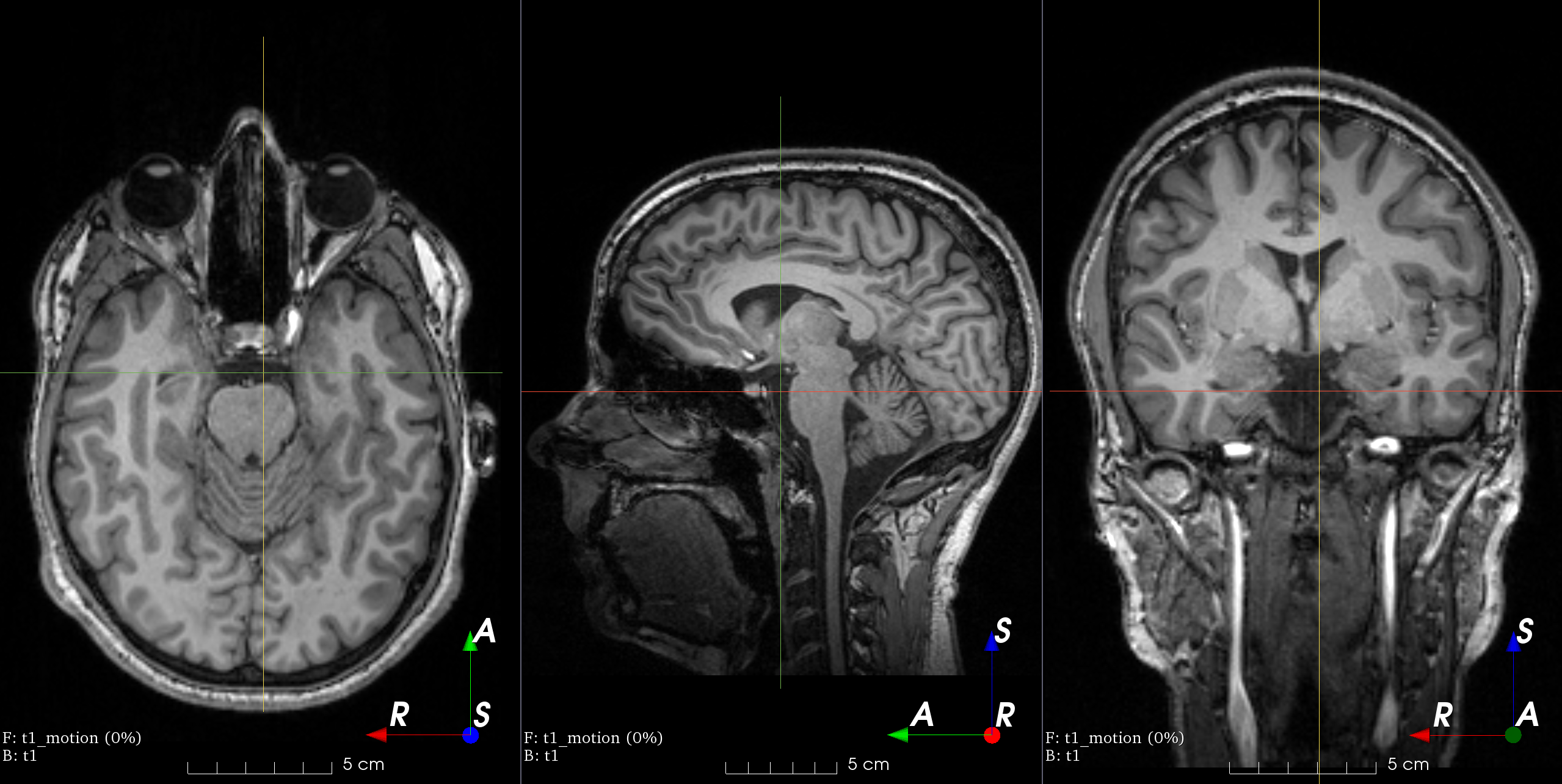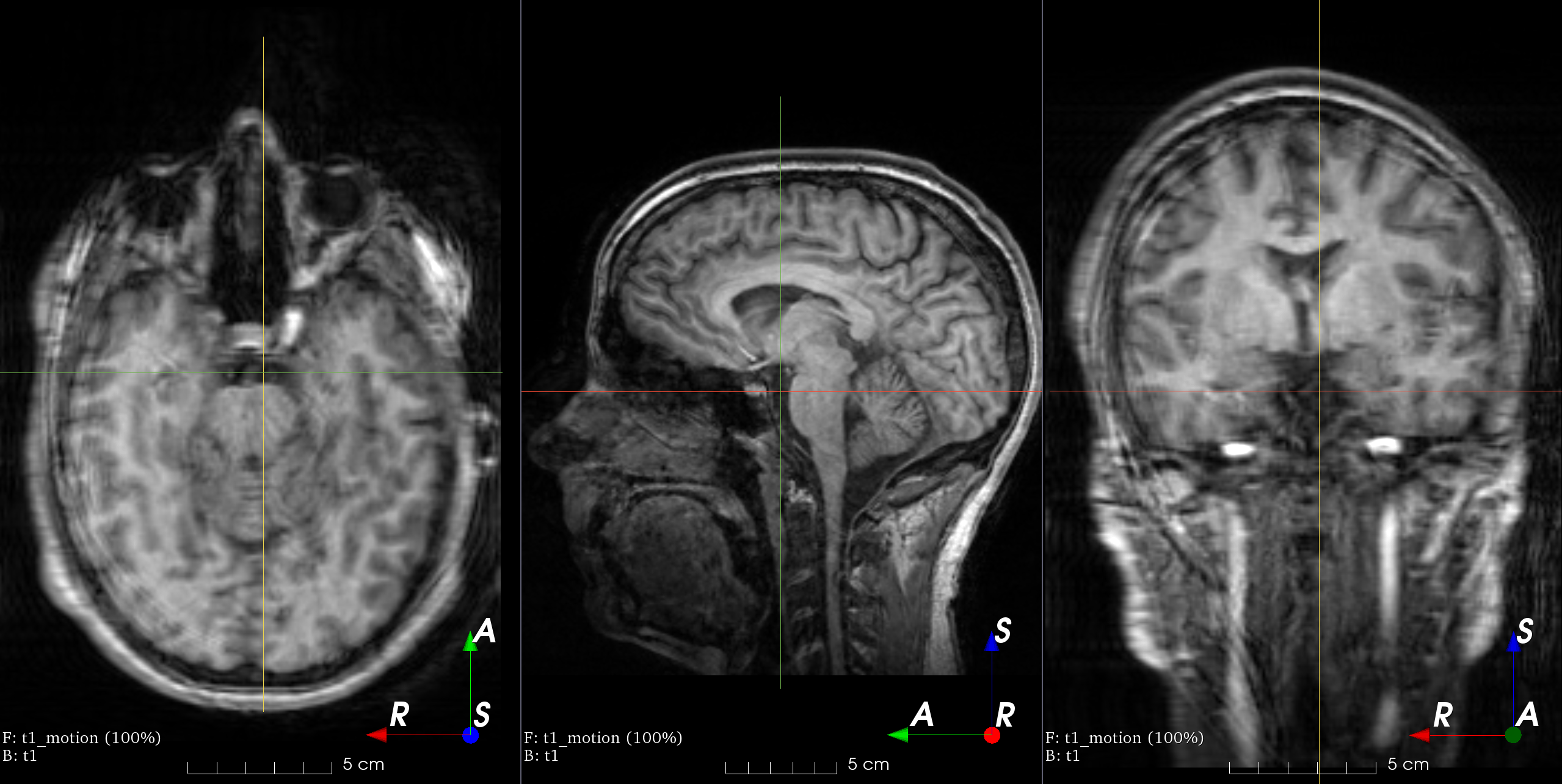
MRI k-space motion artifacts
Magnetic resonance images suffer from motion artifacts when the subject moves
during image acquisition. This transform follows
Shaw et al., 2019 to
simulate motion artifacts for data augmentation.
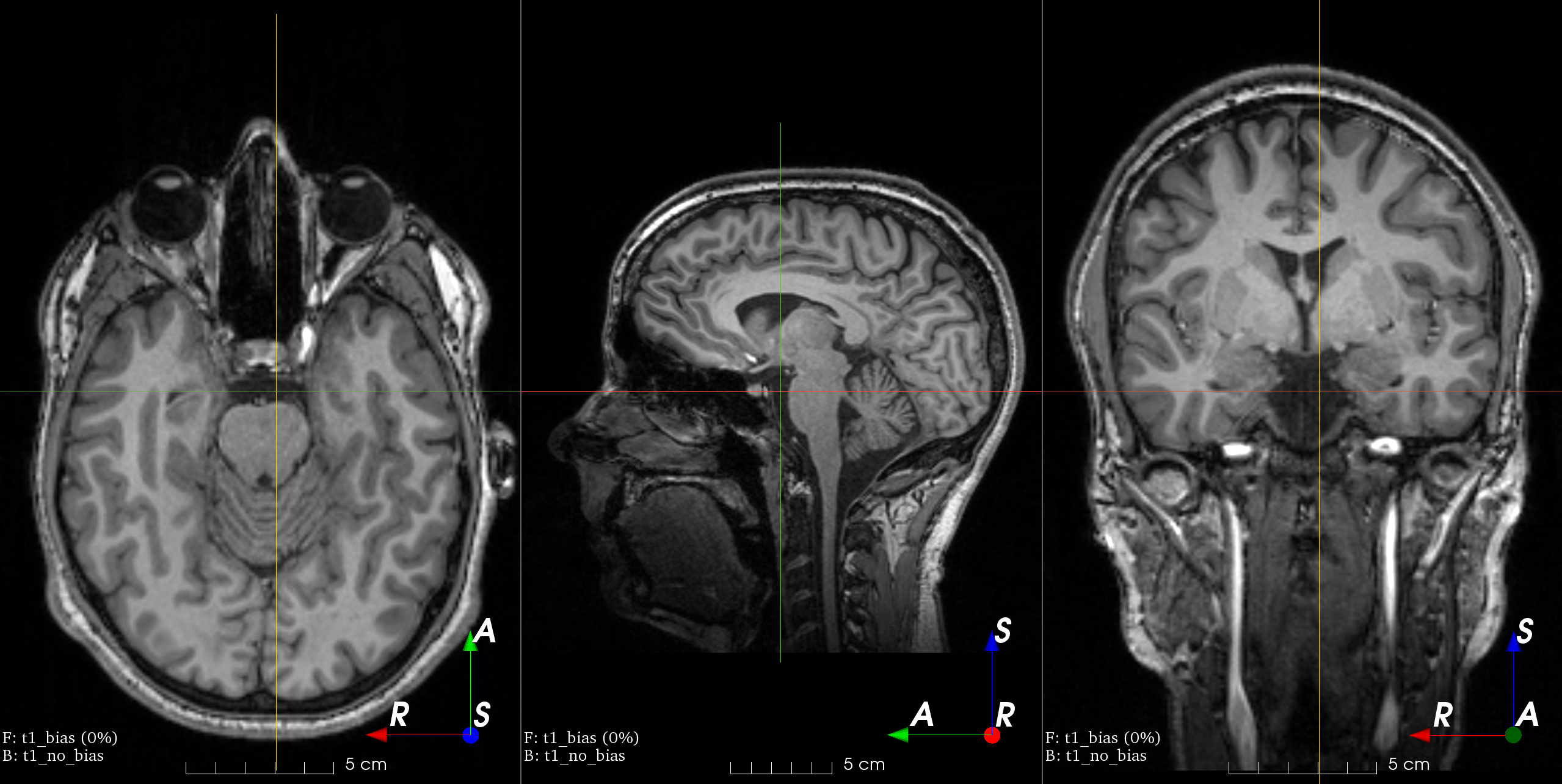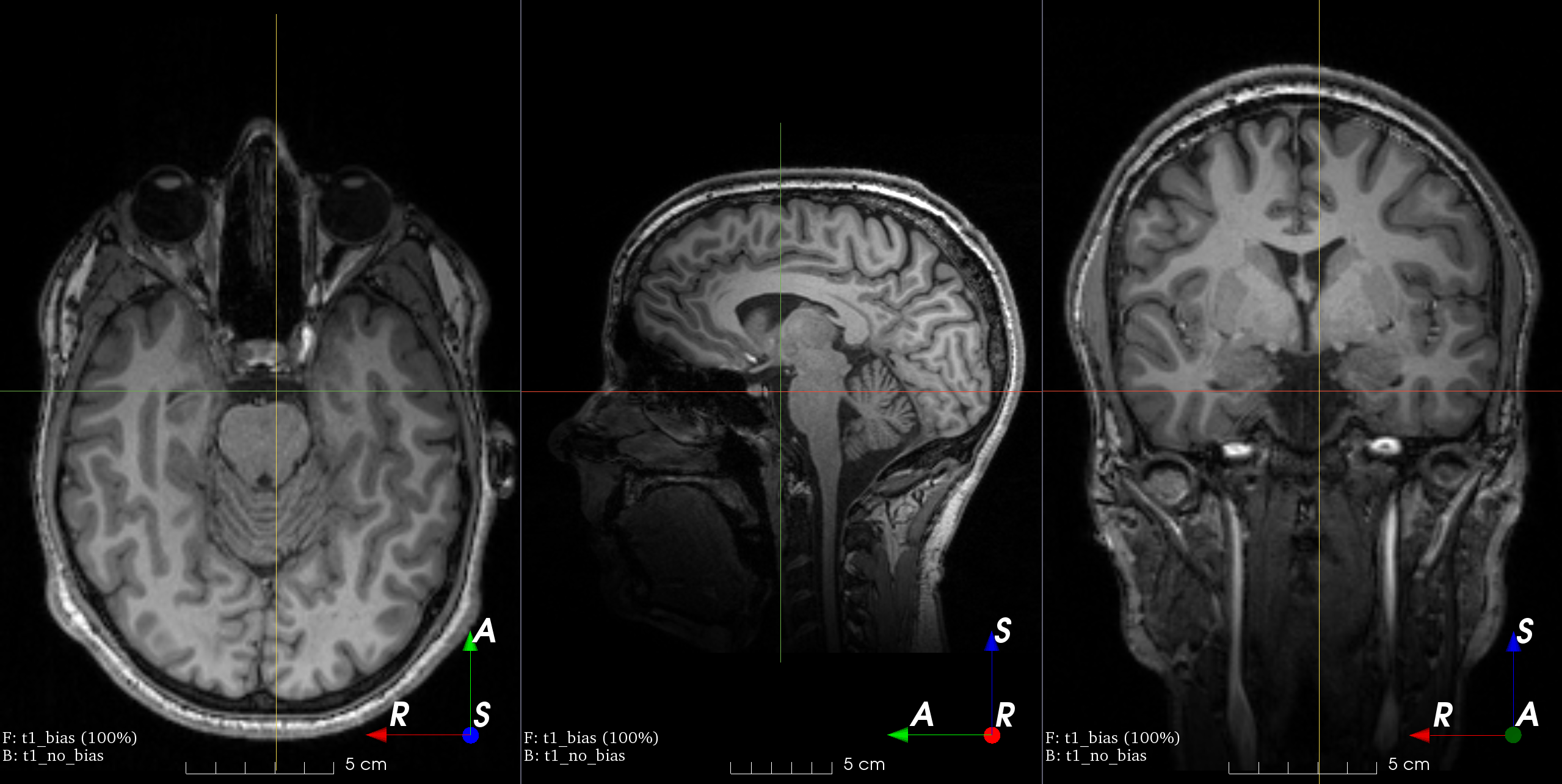
MRI magnetic field inhomogeneity
MRI magnetic field inhomogeneity creates slow frequency intensity variations.
This transform is very similar to the one in
NiftyNet.
Gaussian noise
Adds noise sampled from a normal distribution with mean 0 and standard
deviation sampled from a uniform distribution in the range std_range.
It is often used after ZNormalization, as the output of
this transform has zero-mean.

Normalization
Histogram standardization
Implementation of
New variants of a method of MRI scale standardization
adapted from NiftyNet.
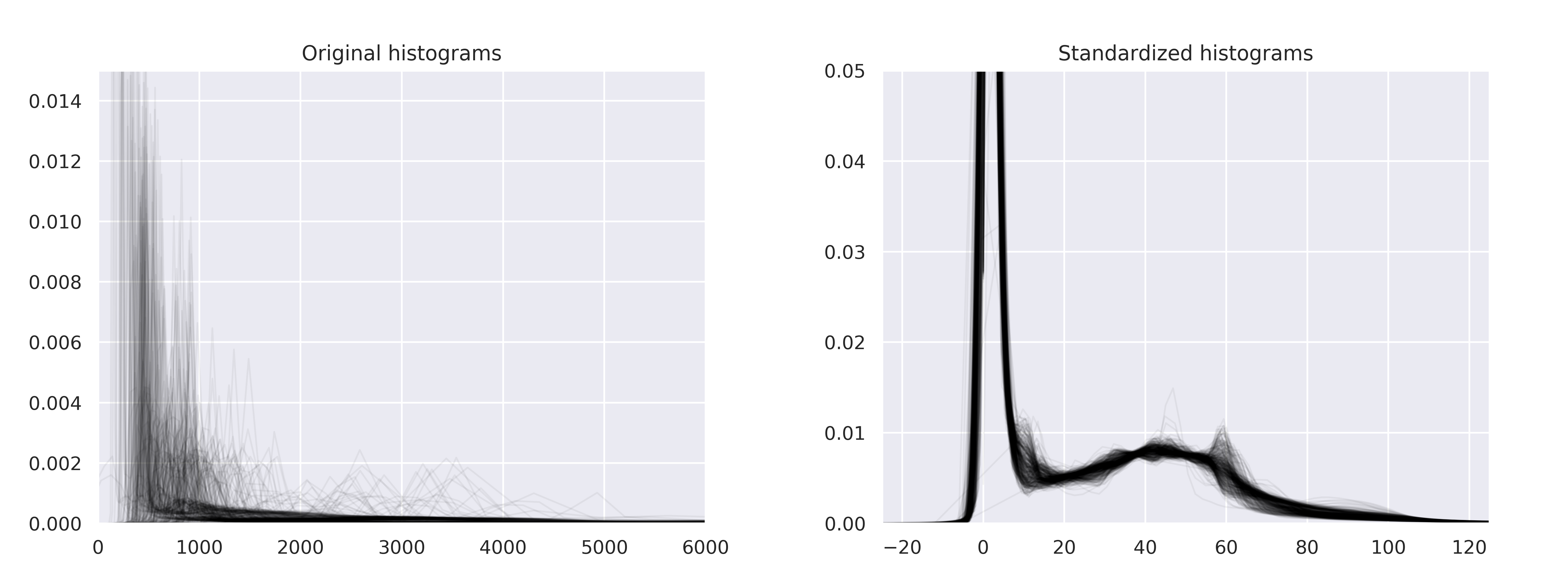
Z-normalization
Rescale
Spatial
Flip
Reverse the order of elements in an image along the given axes.
Affine transform
B-spline dense elastic deformation
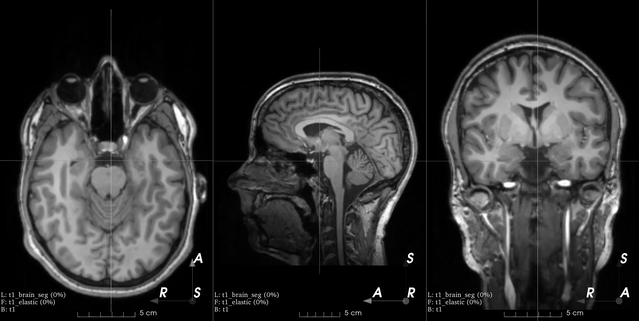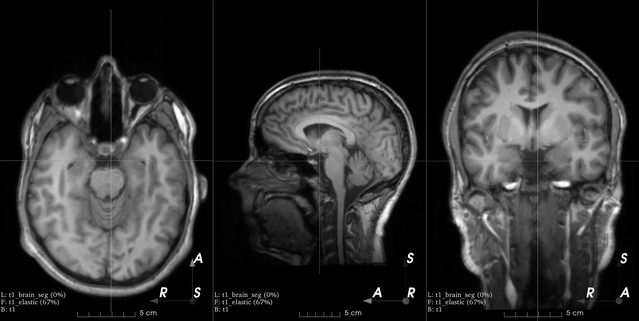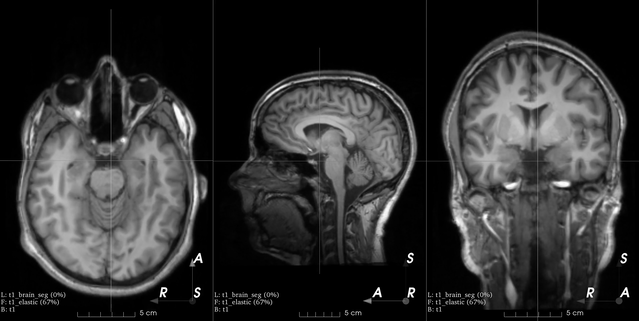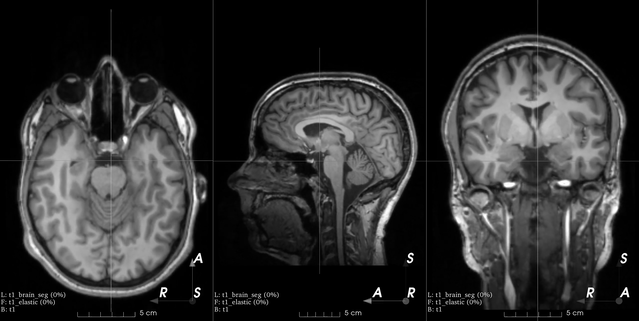
Example
This example shows the improvement in performance when multiple workers are
used to load and preprocess the volumes using multiple workers.
import time
import multiprocessing as mp
from torch.utils.data import DataLoader
from torchvision.transforms import Compose
from torchio import ImagesDataset, Queue
from torchio.sampler import ImageSampler
from torchio.utils import create_dummy_dataset
from torchio.transforms import (
ZNormalization,
RandomNoise,
RandomFlip,
RandomAffine,
)
# Define training and patches sampling parameters
num_epochs = 4
patch_size = 128
queue_length = 100
samples_per_volume = 10
batch_size = 4
def model(batch, sleep_time=0.1):
"""Dummy function to simulate a forward pass through the network"""
time.sleep(sleep_time)
return batch
# Create a dummy dataset in the temporary directory, for this example
subjects_paths = create_dummy_dataset(
num_images=100,
size_range=(193, 229),
force=False,
)
# Each element of subjects_paths is a dictionary:
# subject = {
# 'one_image': dict(path=path_to_one_image, type=torchio.INTENSITY),
# 'another_image': dict(path=path_to_another_image, type=torchio.INTENSITY),
# 'a_label': dict(path=path_to_a_label, type=torchio.LABEL),
# }
# Define transforms for data normalization and augmentation
transforms = (
ZNormalization(),
RandomNoise(std_range=(0, 0.25)),
RandomAffine(scales=(0.9, 1.1), degrees=10),
RandomFlip(axes=(0,)),
)
transform = Compose(transforms)
subjects_dataset = ImagesDataset(subjects_paths, transform)
sample = subjects_dataset[0]
# Run a benchmark for different numbers of workers
workers = range(mp.cpu_count() + 1)
for num_workers in workers:
print('Number of workers:', num_workers)
# Define the dataset as a queue of patches
queue_dataset = Queue(
subjects_dataset,
queue_length,
samples_per_volume,
patch_size,
ImageSampler,
num_workers=num_workers,
)
batch_loader = DataLoader(queue_dataset, batch_size=batch_size)
start = time.time()
for epoch_index in range(num_epochs):
for batch in batch_loader:
logits = model(batch)
print('Time:', int(time.time() - start), 'seconds')
print()
Output:
Number of workers: 0
Time: 394 seconds
Number of workers: 1
Time: 372 seconds
Number of workers: 2
Time: 278 seconds
Number of workers: 3
Time: 259 seconds
Number of workers: 4
Time: 242 seconds



