GNNGuard
GNNGuard: Defending Graph Neural Networks against Adversarial Attacks
Authors: Xiang Zhang ([email protected]), Marinka Zitnik ([email protected])
Overview
This repository contains python codes and datasets necessary to run the GNNGuard algorithm. GNNGuard is a general defense approach against a variety of poisoning adversarial attacks that perturb the discrete graph structure. GNNGuard can be straightforwardly incorporated into any GNN models to prevent the misclassification caused by poisoning adversarial attacks on graphs. Please see our paper for more details on the algorithm.
Key Idea of GNNGuard
Deep learning methods for graphs achieve remarkable performance on many tasks. However, despite the proliferation of such methods and their success, recent findings indicate that small, unnoticeable perturbations of graph structure can catastrophically reduce performance of even the strongest and most popular Graph Neural Networks (GNNs). By integrating with the proposed GNNGuard, the GNN classifier can correctly classify the target node even under strong adversarial attacks.
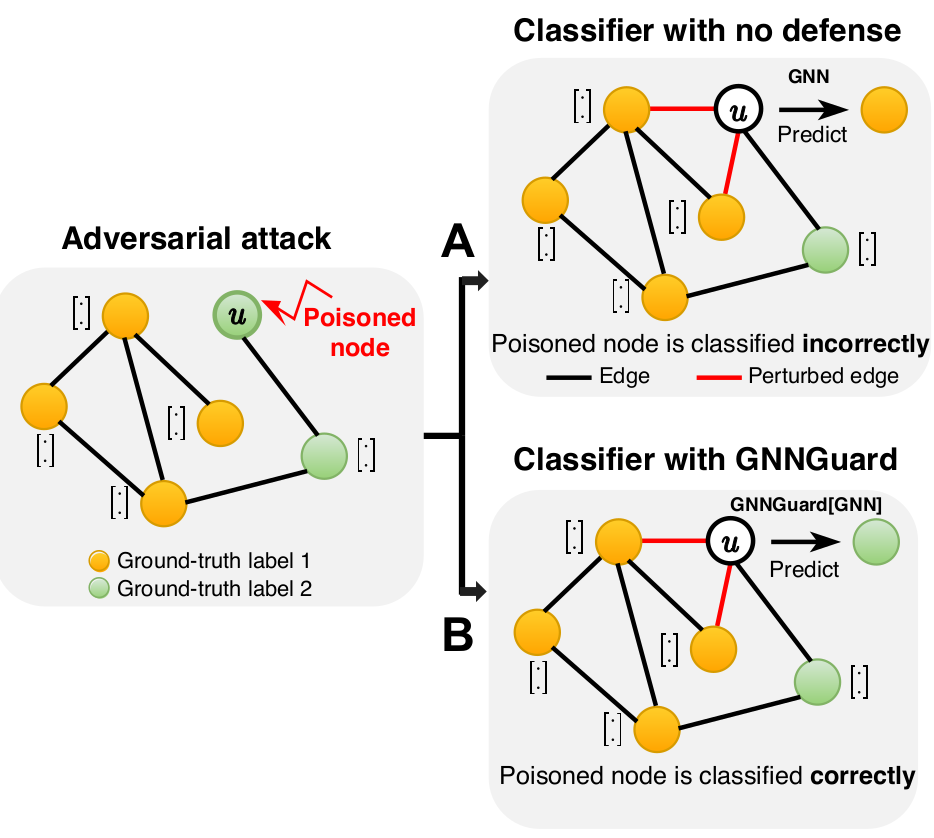
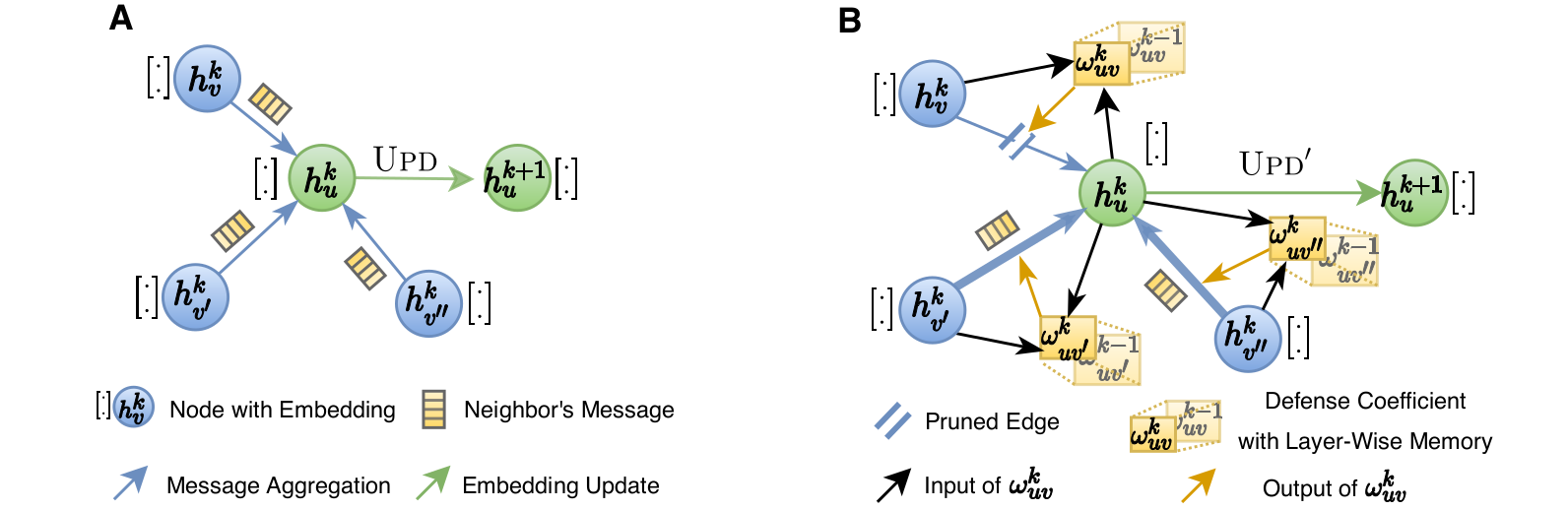
The key idea of GNNGuard is to detect and quantify the relationship between the graph structure and node features, if one exists, and then exploit that relationship to mitigate negative effects of the attack. GNNGuard learns how to best assign higher weights to edges connecting similar nodes while pruning edges between unrelated nodes. In specific, instead of the neural message passing of typical GNN (shown as A), GNNGuard (B) controls the message stream such as blocking the message from irrelevent neighbors but strengthening messages from highly-related ones. Importantly, we are the first model that can defend heterophily graphs (\eg, with structural equivalence) while all the existing defenders only considering homophily graphs.
Running the code
The GNNGuard is evluated under three typical adversarial attacks including Direct Targeted Attack (Nettack-Di), Influence Targeted Attack (Nettack-In), and Non-Targeted Attack (Mettack). In GNNGuard folder, the Nettack-Di.py, Nettack-In.py, and Mettack.py corresponding to the three adversarial attacks.
For example, to check the performance of GCN without defense under direct targeted attack, run the following code:
python Nettack-Di.py --dataset Cora --modelname GCN --GNNGuard False
Turn on the GNNGuard defense, run
python Nettack-Di.py --dataset Cora --modelname GCN --GNNGuard True
Note: Please uncomment the defense models (Line 144 for Nettack-Di.py) to test different defense models.
Citing
If you find GNNGuard useful for your research, please consider citing this paper:
@inproceedings{zhang2020gnnguard,
title = {GNNGuard: Defending Graph Neural Networks against Adversarial Attacks},
author = {Zhang, Xiang and Zitnik, Marinka},
booktitle = {NeurIPS},
year = {2020}
}
Requirements
GNNGuard is tested to work under Python >=3.5.
Recent versions of Pytorch, torch-geometric, numpy, and scipy are required. All the required basic packages can be installed using the following command:
'''
pip install -r requirements.txt
'''
Note: For toch-geometric and the related dependices (e.g., cluster, scatter, sparse), the higher version may work but haven't been tested yet.
Install DeepRobust
During the evaluation, the adversarial attacks on graph are performed by DeepRobust from MSU, please install it by
git clone https://github.com/DSE-MSU/DeepRobust.git
cd DeepRobust
python setup.py install
- If you have trouble in installing DeepRobust, please try to replace the provided 'defense/setup.py' to replace the original
DeepRobust-master/setup.pyand manully reinstall it by
python setup.py install
-
We extend the original DeepRobust from single GCN to multiplye GNN variants including GAT, GIN, Jumping Knowledge, and GCN-SAINT. After installing DeepRobust, please replace the origininal folder
DeepRobust-master/deeprobust/graph/defenseby thedefensefolder that provided in our repository! -
To better plugin GNNGuard to geometric codes, we slightly revised some functions in geometric. Please use the three files under our provided
nn/conv/to replace the corresponding files in the installed geometric folder (for example, the folder path could be/home/username/.local/lib/python3.5/site-packages/torch_geometric/nn/conv/).
Note: 1). Don't forget to backup all the original files when you replacing anything, in case you need them at other places! 2). Please install the corresponding CUDA versions if you are using GPU.
Datasets
Here we provide the datasets (including Cora, Citeseer, ogbn-arxiv, and DP) used in GNNGuard paper.
The ogbn-arxiv dataset can be easily access by python codes:
from ogb.nodeproppred import PygNodePropPredDataset
dataset = PygNodePropPredDataset(name = 'ogbn-arxiv')
More details about ogbn-arxiv dataset can be found here.
Find more details about Disease Pathway dataset at here.
For graphs with structural roles, a prominent type of heterophily, we calculate the nodes' similarity using graphlet degree vector instead of node embedding. The graphlet degree vector is generated/counted based on the Orbit Counting Algorithm (Orca).



