Graphormer
By Chengxuan Ying, Tianle Cai, Shengjie Luo, Shuxin Zheng*, Guolin Ke, Di He*, Yanming Shen and Tie-Yan Liu.
This repo is the official implementation of "Do Transformers Really Perform Bad for Graph Representation?".
Introduction
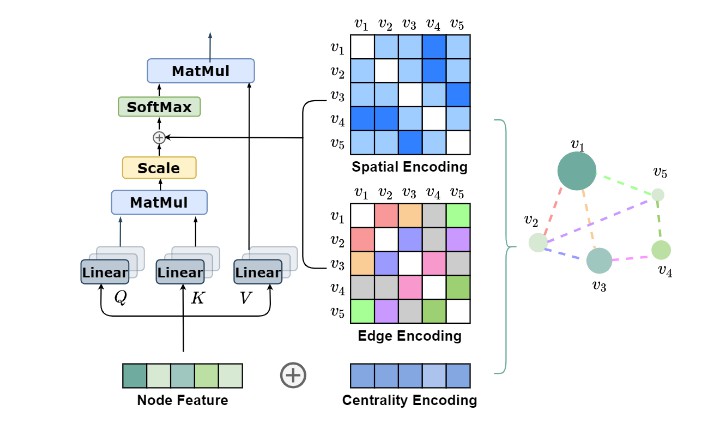
Graphormer is initially described in arxiv, which is a standard Transformer architecture with several structural encodings, which could effectively encoding the structural information of a graph into the model.
Graphormer achieves strong performance on PCQM4M-LSC (0.1234 MAE on val), MolPCBA (31.39 AP(%) on test), MolHIV (80.51 AUC(%) on test) and ZINC (0.122 MAE on test), surpassing previous models by a large margin.
Main Results
PCQM4M-LSC
| Method | #params | train MAE | valid MAE |
|---|---|---|---|
| GCN | 2.0M | 0.1318 | 0.1691 |
| GIN | 3.8M | 0.1203 | 0.1537 |
| GCN-VN | 4.9M | 0.1225 | 0.1485 |
| GIN-VN | 6.7M | 0.1150 | 0.1395 |
| Graphormer-Small | 12.5M | 0.0778 | 0.1264 |
| Graphormer | 47.1M | 0.0582 | 0.1234 |
OGBG-MolPCBA
| Method | #params | test AP (%) |
|---|---|---|
| DeeperGCN-VN+FLAG | 5.6M | 28.42 |
| DGN | 6.7M | 28.85 |
| GINE-VN | 6.1M | 29.17 |
| PHC-GNN | 1.7M | 29.47 |
| GINE-APPNP | 6.1M | 29.79 |
| Graphormer | 119.5M | 31.39 |
OGBG-MolHIV
| Method | #params | test AP (%) |
|---|---|---|
| GCN-GraphNorm | 526K | 78.83 |
| PNA | 326K | 79.05 |
| PHC-GNN | 111K | 79.34 |
| DeeperGCN-FLAG | 532K | 79.42 |
| DGN | 114K | 79.70 |
| Graphormer | 47.0M | 80.51 |
ZINC-500K
| Method | #params | test MAE |
|---|---|---|
| GIN | 509.5K | 0.526 |
| GraphSage | 505.3K | 0.398 |
| GAT | 531.3K | 0.384 |
| GCN | 505.1K | 0.367 |
| GT | 588.9K | 0.226 |
| GatedGCN-PE | 505.0K | 0.214 |
| MPNN (sum) | 480.8K | 0.145 |
| PNA | 387.2K | 0.142 |
| SAN | 508.6K | 0.139 |
| Graphormer-Slim | 489.3K | 0.122 |
Requirements and Installation
Setup with Conda
# create a new environment
conda create --name graphormer python=3.7
conda activate graphormer
# install requirements
pip install rdkit-pypi cython
pip install ogb==1.3.1 pytorch-lightning==1.3.0
pip install torch==1.7.1+cu110 torchvision==0.8.2+cu110 -f https://download.pytorch.org/whl/torch_stable.html
pip install torch-geometric==1.6.3 ogb==1.3.1 pytorch-lightning==1.3.1 tqdm torch-sparse==0.6.9 torch-scatter==2.0.6 -f https://pytorch-geometric.com/whl/torch-1.7.0+cu110.html
Citation
Please kindly cite this paper if you use the code:
@article{ying2021transformers,
title={Do Transformers Really Perform Bad for Graph Representation?},
author={Ying, Chengxuan and Cai, Tianle and Luo, Shengjie and Zheng, Shuxin and Ke, Guolin and He, Di and Shen, Yanming and Liu, Tie-Yan},
journal={arXiv preprint arXiv:2106.05234},
year={2021}
}






