ngstools
My own tools code for NGS data analysis (Next Generation Sequencing)
Now we have 2 R functions and 1 Python tool:
Python tools:
parse-mpileup.py
R functions:
plot.TAD
plot.matrix
How to cite?
When you use code from this repository, please cite like:
MENG Haowei, https://github.com/menghaowei/ngstools
R Function: plot.matrix()
please download ngs_R_function.R, ./test_data/hic-chr_1_100000.matrix.txt" and you can use those R functions directly with source command in R envirnment like:
# load R funtions
source(file = "ngs_R_function.R")
# load test matrix
# This test matrix is from a real Hi-C data set
test_mat = read.table(file = "./test_data/hic-chr_1_100000.matrix.txt",header = F,sep = ",")
1.plot matrix
plot.matrix(test_mat)
title(main="plot heatmap from Hi-C data")

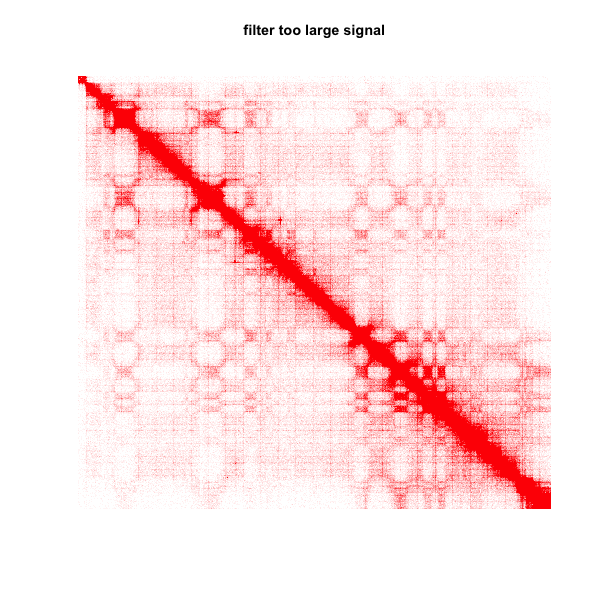
2.plot matrix and filter too large signal (95% quantile number as cutoff)
plot.matrix(test_mat,bound.max = 0.95)
title(main="filter too large signal")
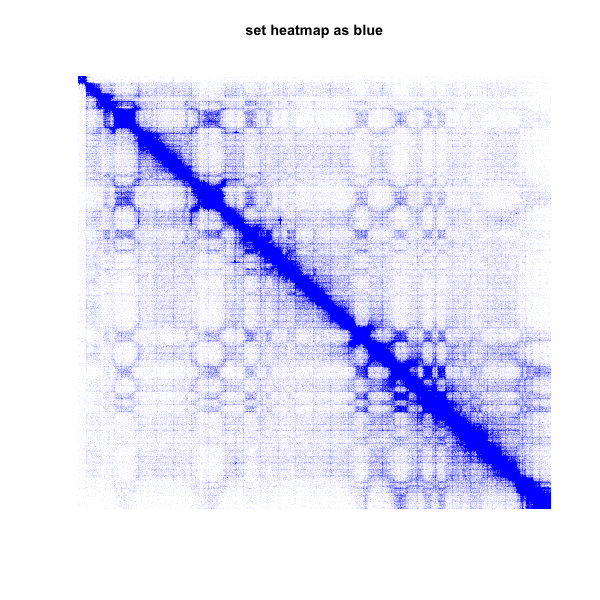
3.set color range from blue to blue
plot.matrix(test_mat,bound.max = 0.95,col.min = "blue",col.max = "blue")
title(main="set heatmap as blue")
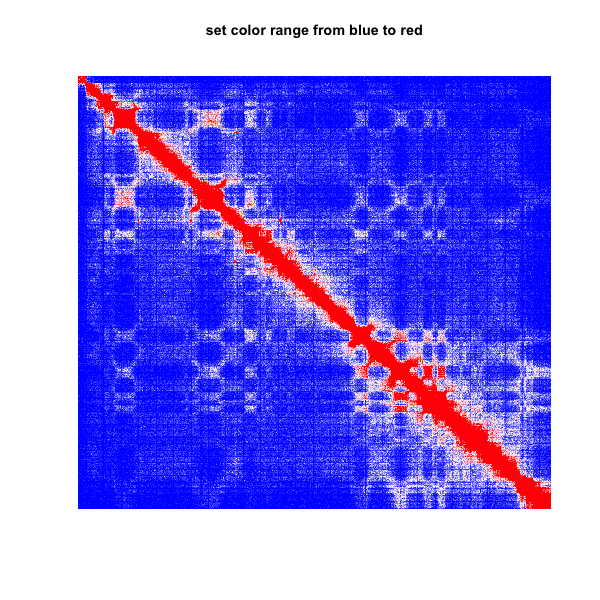
4.set color range from blue to red (the bins number less 10 are set as blue)
plot.matrix(test_mat,bound.max = 0.95,col.min = "blue",col.max = "red",col.boundary = 10)
title(main="set color range from blue to red")
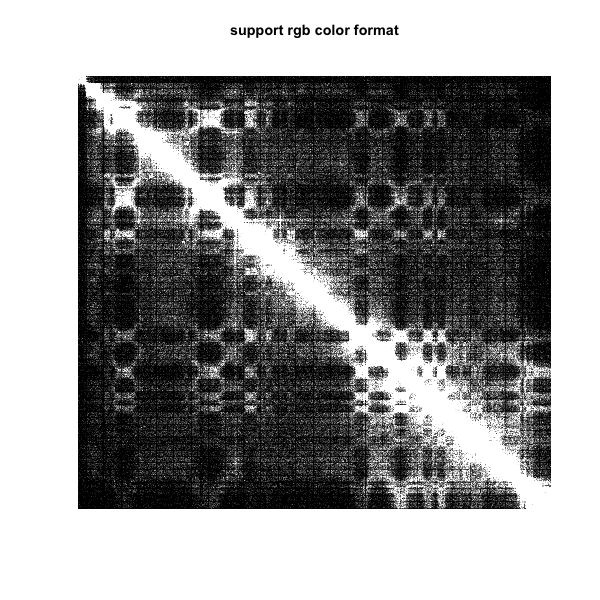
5.the function also support RGB color, #000000 means black and #FFFFFF means white
plot.matrix(test_mat,bound.max = 0.95,col.min = "#000000",col.max = "#FFFFFF",col.boundary = 10)
title(main="support rgb color format")

R Function: plot.TAD()
please download ngs_R_function.R, ./test_data/hic-chr_1_100000.matrix.txt" and you can use those R functions directly with source command in R envirnment like:
# load R funtions
source(file = "ngs_R_function.R")
# load test matrix
# This test matrix is from a real Hi-C data set
test_mat = read.table(file = "./test_data/hic-chr_1_100000.matrix.txt",header = F,sep = ",")
1.plot upper triangle of matrix (useful for Hi-C TAD plot)
plot.TAD(test_mat,maxBound = 0.95)
title(main="plot TAD with the same Hi-C matrix")
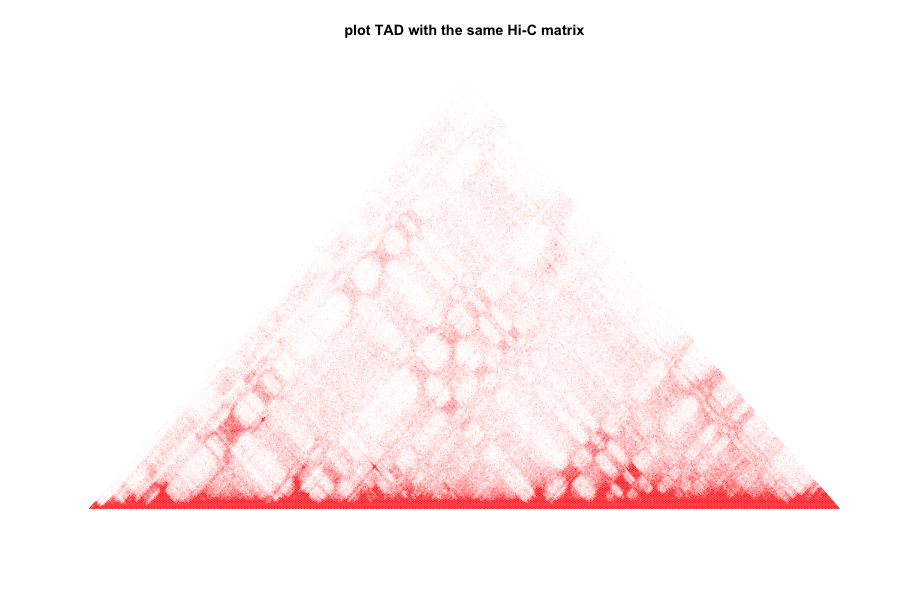

2.plot lower triangle of matrix
plot.TAD(test_mat,maxBound = 0.95,mat.upper = F)
title(main="plot lower triangle of matrix")

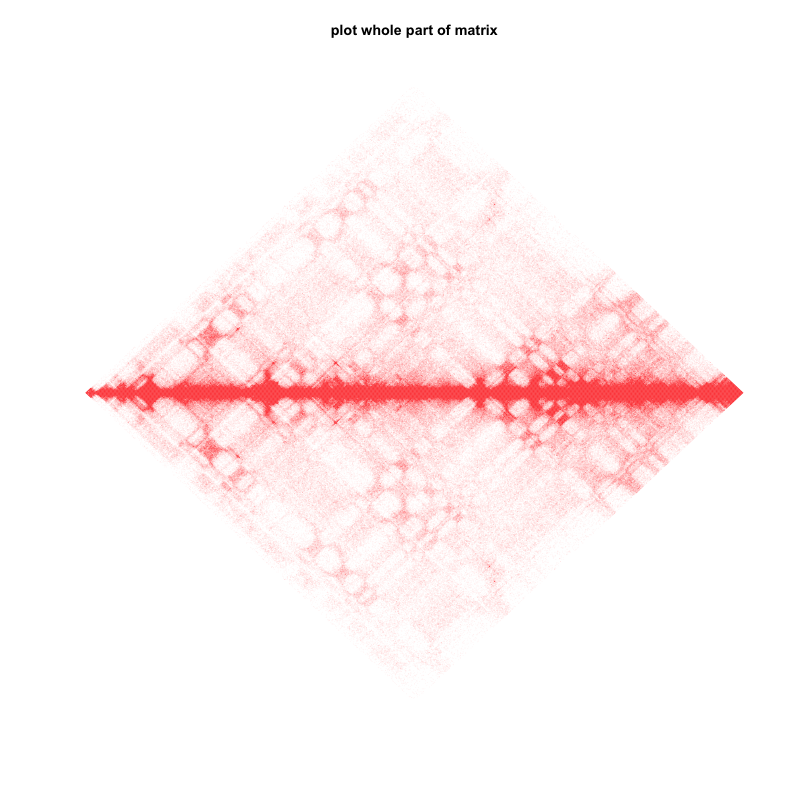
3.plot all matrix
plot.TAD(test_mat,maxBound = 0.95,mat.part = F)
title(main="plot whole part of matrix")
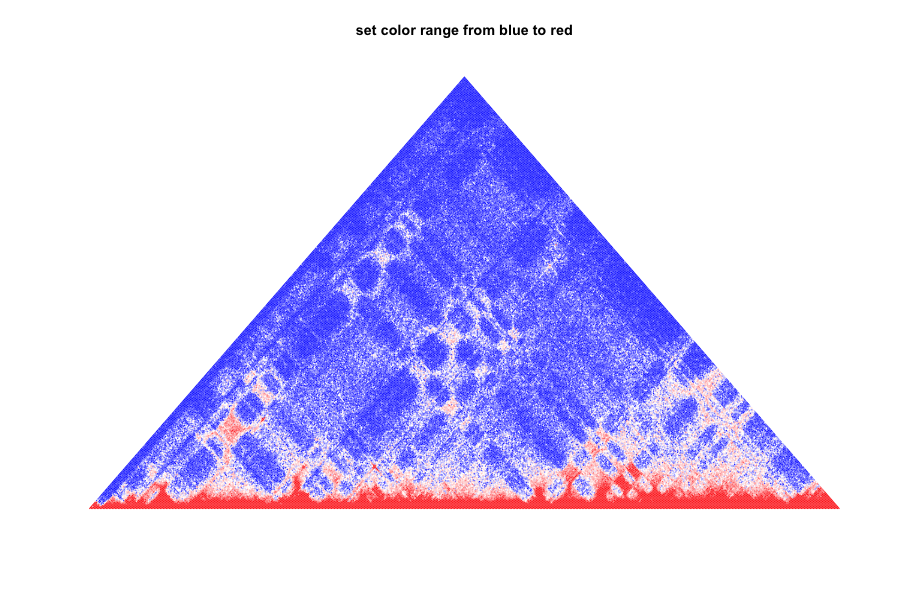
4.set color range from blue to red (the bins number less 10 are set as blue)
plot.TAD(test_mat,maxBound = 0.95,col.min = "blue",col.max = "red",col.boundary = 5)
title(main="set color range from blue to red")
parse-mpileup.py
parse samtools mpileup command output, also known as .pileup file, the file like:
chr1 10030 c 1 ^6. A
chr1 10031 t 1 . A
chr1 10032 a 1 . F
chr1 10033 a 1 . F
chr1 10034 c 1 . <
chr1 10035 c 1 . F
chr1 10036 c 0 * *
chr1 10037 t 1 .-1A A
chr1 10038 a 0 * *
chr1 10039 a 0 * *
The .pileup format explain please check the HTML
pileup explain
And convert .pileup file into .bmatformat, the format like:
chr_name chr_index ref_base A G C T del_count insert_count ambiguous_count deletioninsertion ambiguous mut_num
chr1 10030 C 0 0 1 0 0 0 0 . . . 0
chr1 10031 T 0 0 0 1 0 0 0 . . . 0
chr1 10032 A 1 0 0 0 0 0 0 . . . 0
chr1 10033 A 1 0 0 0 0 0 0 . . . 0
chr1 10034 C 0 0 1 0 0 0 0 . . . 0
chr1 10035 C 0 0 1 0 0 0 0 . . . 0
chr1 10036 C 0 0 0 0 1 0 0 * . . 0
chr1 10037 T 0 0 0 1 1 0 0 A . . 0
chr1 10038 A 0 0 0 0 1 0 0 * . . 0
For help info, please run python parse-mpileup.py -h:
python parse-mpileup.py -h
usage: parse-mpileup.py [-h] -i INPUT [-o OUTPUT] [-p THREADS] [-n MUTNUM]
[--TempDir TEMPDIR]
convert mpileup file to info file
optional arguments:
-h, --help show this help message and exit
-i INPUT, --Input INPUT
samtools mpileup format file
-o OUTPUT, --Output OUTPUT
Output parsed file
-p THREADS, --Threads THREADS
Multiple threads number, default=1
-n MUTNUM, --MutNum MUTNUM
Only contain mutation info go to the output, set 0
mean output all site, default=0
--TempDir TEMPDIR Where to keep temp files, default is the same dir with
--Input




